Cancer is a complex disease that results from a combination of genetic and environmental perturbations to biomolecular networks that, in healthy tissues, maintain a homeostatic balance between normal cellular functional states. Computational and statistical techniques for analyzing primary tumors, metastases, or cell culture systems can help us to understand cellular dysfunction underlying cancer onset, progression, and metastasis as well as to develop novel diagnostic and prognostic tools.
B. Tercan, G. Qin, T-K. Kim, B. Aguilar, J. Phan, W. Longabaugh, D. Pot, C. J. Kemp, N. Chambwe, I. Shmulevich, “SL-Cloud: A Cloud-based resource to support synthetic lethal interaction discovery,” F1000Research, Vol. 11, No. 493, 2022.
R. Moser, K. E. Gurley, O. Nikolova, G. Qin, R. Joshi, E. Mendez, I. Shmulevich, A. Ashley, C. Grandori, C. J. Kemp, “Synthetic lethal kinases in Ras/p53 mutant squamous cell carcinoma,” Oncogene, Vol 41, No. 24, pp. 3355-3369, 2022.
E. A. Stahlberg, M. Abdel-Rahman, B. Aguilar, A. Asadpoure, R. A. Beckman, L. L. Borkon, J. Bryan, C. Cebulla, Y. H. Chang, A. Chatterjee, J. Deng, S. Dolatshahi, O. Gevaert, E. J. Greenspan, W. Hao, T. Hernandez-Boussard, P. R. Jackson, M. Kuijjer, A. Lee, P. Macklin, S. Madhavan, M. D. McCoy, N. M. Mirzaei, T. Razzaghi, H. Rocha, L. Shahriyari, I. Shmulevich, D. G. Stover, Y. Sun, T. Syeda-Mahmood, J. Wang, Q. Wang, and I. Zervantonakis, “Exploring approaches for Predictive cancer patient digital twins: Opportunities for collaboration and Innovation,” Frontiers in Digital Health, vol. 4, 2022.
G. Qin, T. A. Knijnenburg, D. L. Gibbs, R. Moser, R. J. Monnat Jr., C. J. Kemp, I. Shmulevich, “A functional module states framework reveals transcriptional states for drug and target prediction,” Cell Reports, Vol. 38, No. 3, 110269, 2022.
T. Hernandez-Boussard, P. Macklin, E. J. Greenspan, A. L. Gryshuk, E. Stahlberg, T. Syeda-Mahmood, I. Shmulevich, “Digital Twins for predictive oncology will be a paradigm shift for precision cancer care,” Nature Medicine, (PMID: 34824458), 2021.
D. L. Gibbs, B. Aguilar, V. Thorsson, A. V. Ratushny, I. Shmulevich, “Patient-specific cell communication networks associate with disease progression in cancer,” Frontiers in Genetics, Vol. 12, 667382, 2021.
Y. H. Lo, K. Kolahi, Y. Du, C. Y. Chang, A. Krokhotin, A. Nair, W. Sobba, K. Karlsson, S. Jones, T. Longacre, A. Mah, B. Tercan, A. Sockell, H. Xu, J. Seoane, J. Chen, I. Shmulevich, J. Weissman, C. Curtis, A. Califano, H. Fu, G. Crabtree, C. J. Kuo, “A CRISPR/Cas9-engineered ARID1A-deficient human gastric cancer organoid model reveals essential and non-essential modes of oncogenic transformation,” Cancer Discovery, Vol. 11, No. 6, pp. 1562-1581, 2021.
S. Cao, D. C. Zhou, C. Oh, R. G. Jayasinghe, Y. Zhao, C. J. Yoon, M. A. Wyczalkowski, M. H. Bailey, T. Tsou, Q. Gao, A. Malone, S. Reynolds, I. Shmulevich, M. C. Wendl, F. Chen, L. Ding, “Discovery of driver non-coding splice-site-creating mutations in cancer,” Nature Communications, Vol. 11, No. 1, 5573, 2020.
ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium, “Pan-cancer analysis of whole genomes,” Nature, Vol. 578, No. 7793, pp. 82-93, 2020.
S. Cao, D. C. Zhou, C. Oh, R. G. Jayasinghe, Y. Zhao, C. J. Yoon, M. A. Wyczalkowski, M. H. Bailey, T. Tsou, Q. Gao, A. Malone, S. Reynolds, I. Shmulevich, M. C. Wendl, F. Chen, L. Ding, “Discovery of driver non-coding splice-site-creating mutations in cancer,” Nature Communications, Vol. 11, No. 1, 5573, 2020.
J. A. Eddy, V. Thorsson, A. E. Lamb, D. L. Gibbs, C. Heimann, J. X. Yu, V. Chung, Y. Chae, K. Dang, B. G. Vincent, I. Shmulevich, J. Guinney, “CRI iAtlas: an interactive portal for immuno-oncology research,” F1000Research, Vol. 9, 1028, 2020.
J. B. Xavier, V. B. Young, J. Skufca, F. Ginty, T. Testerman, A. T. Pearson, P. Macklin, A. Mitchell, I. Shmulevich, L. Xie, J. G. Caporaso, K. A. Crandall, N. L. Simone, F. Godoy-Vitorino, T. J. Griffin, K. L. Whiteson, H. H. Gustafson, D. J. Slade, T. M. Schmidt, M. R. S. Walther-Antonio, T. Korem, B. M. Webb-Robertson, M. P. Styczynski, W. E. Johnson, C. Jobin, J. M. Ridlon, A. Y. Koh, M. Yu, L. Kelly, J. A. Wargo, “The Cancer Microbiome: Distinguishing Direct and Indirect Effects Requires a Systemic View,” Trends in Cancer, Vol. 6, No. 3, pp. 192-204, 2020.
J. Orozco, T. Knijnenburg, A. Manughian-Peter, M. Salomon, G. Barkhoudarian, J. Jalas, J. Wilmott, P. Hothi, X. Wang, Y. Takasumi, M. Buckland, J. Thompson, G. Long, C. Cobbs, I. Shmulevich, D. Kelly, R. Scolyer, D. Hoon, D. Marzese, “Epigenetic profiling for the molecular classification of metastatic brain tumors,” Nature Communications, Vol. 9, No. 4627, 2018.
V. Thorsson, D. Gibbs, S. Brown, D. Wolf, D. Bortone, T. Ou Yang, E. Porta-Pardo, G. Gao, C. Plaisier, J. Eddy, E. Ziv, A. Culhane, E. Paull, I. Sivakumar, A. Gentles, R. Malhotra, F. Farshidfar, A. Colaprico, J. Parker, L. Mose, N. Vo, J. Liu, Y. Liu, J. Rader, V. Dhankani, S. Reynolds, R. Bowlby, A. Califano, A. Cherniack, D. Anastassiou, D. Bedognetti, A. Rao, K. Chen, A. Krasnitz, H. Hu, T. Malta, H. Noushmehr, C. Pedamallu, S. Bullman, A. Ojesina, A. Lamb, W. Zhou, H. Shen, T. Choueiri, J. Weinstein, J. Guinney, J. Saltz, R. Holt, C. Rabkin, Cancer Genome Atlas Research Network, A. Lazar, J. Serody, E. Demicco, M. Disis, B. Vincent, l. Shmulevich, “The Immune Landscape of Cancer“, Immunity, Vol. 48, No. 4, pp. 812-830.e14, 2018.
K. Huang, R. Mashl, Y. Wu, D. Ritter, J. Wang, C. Oh, M. Paczkowska, S. Reynolds, M. Wyczalkowski, N. Oak, A. Scott, M. Krassowski, A. Cherniack, K. Houlahan, R. Jayasinghe, L. Wang, D. Zhou, D. Liu, S. Cao, Y. Kim, A. Koire, J. McMichael, V. Hucthagowder, T. Kim, A. Hahn, C. Wang, M. McLellan, F. Al-Mulla, K. Johnson, O. Cancer Genome Atlas Research Network, Lichtarge, P. Boutros, B. Raphael, A. Lazar, W. Zhang, M. Wendl, R. Govindan, S. Jain, D. Wheeler, S. Kulkarni, J. Dipersio, J. Reimand, F. Meric-Bernstam, K. Chen, I. Shmulevich, S. Plon, F. Chen, L. Ding, “Pathogenic Germline Variants in 10,389 Adult Cancers“, Cell, vol. 173, no. 2, pp. 355-370.e14, 2018.
F. Sanchez-Vega, M. Mina, J. Armenia, W. Chatila, A. Luna, K. La, S. Dimitriadoy, D. Liu, H. Kantheti, S. Saghafinia, D. Chakravarty, F. Daian, Q. Gao, M. Bailey, W. Liang, S. Foltz, I. Shmulevich, L. Ding, Z. Heins, A. Ochoa, B. Gross, J. Gao, H. Zhang, R. Kundra, C. Kandoth, I. Bahceci, L. Dervishi, U. Dogrusoz, W. Zhou, H. Shen, P. Laird, G. Way, C. Greene, H. Liang, Y. Xiao, C. Wang, A. Iavarone, A. Berger, T. Bivona, A. Lazar, G. Hammer, T. Giordano, L. Kwong, G. McArthur, C. Huang, A. Tward, M. Frederick, F. McCormick, M. Meyerson, Cancer Genome Atlas Research Network, E. Van Allen, A. Cherniack, G. Ciriello, C. Sander, N. Schultz, S. “Oncogenic Signaling Pathways in The Cancer Genome Atlas“, Cell, Vol. 173, No. 2, pp. 321-337.e10, 2018.
L. Ding, M. Bailey, E. Porta-Pardo, V. Thorsson, A. Colaprico, D. Bertrand, D. Gibbs, A. Weerasinghe, K. Huang, C. Tokheim, I. Cortés-Ciriano, R. Jayasinghe, F. Chen, L. Yu, S. Sun, C. Olsen, J. Kim, A. Taylor, A. Cherniack, R. Akbani, C. Suphavilai, N. Nagarajan, J. Stuart, G. Mills, M. Wyczalkowski, B. Vincent, C. Hutter, J. Zenklusen, K. Hoadley, M. Wendl, l. Shmulevich, A. Lazar, D. Wheeler, G. Getz, Cancer Genome Atlas Research Network, “Perspective on Oncogenic Processes at the End of the Beginning of Cancer Genomics“, Cell, Vol. 173, No. 2, pp. 305-320.e10, 2018.
R. Jayasinghe, S. Cao, Q. Gao, M. Wendl, N. Vo, S. Reynolds, Y. Zhao, H. Climente-González, S. Chai, F. Wang, R. Varghese, M. Huang, W. Liang, M. Wyczalkowski, S. Sengupta, Z. Li, S. Payne, D. Fenyö, J. Miner, M. Walter, Cancer Genome Atlas Research Network, B. Vincent, E. Eyras, K. Chen, I. Shmulevich, F. Chen, L. Ding, “Systematic Analysis of Splice-Site-Creating Mutations in Cancer“, Cell Reports, Vol. 23, No. 1, pp. 270-281.e3, 2018.
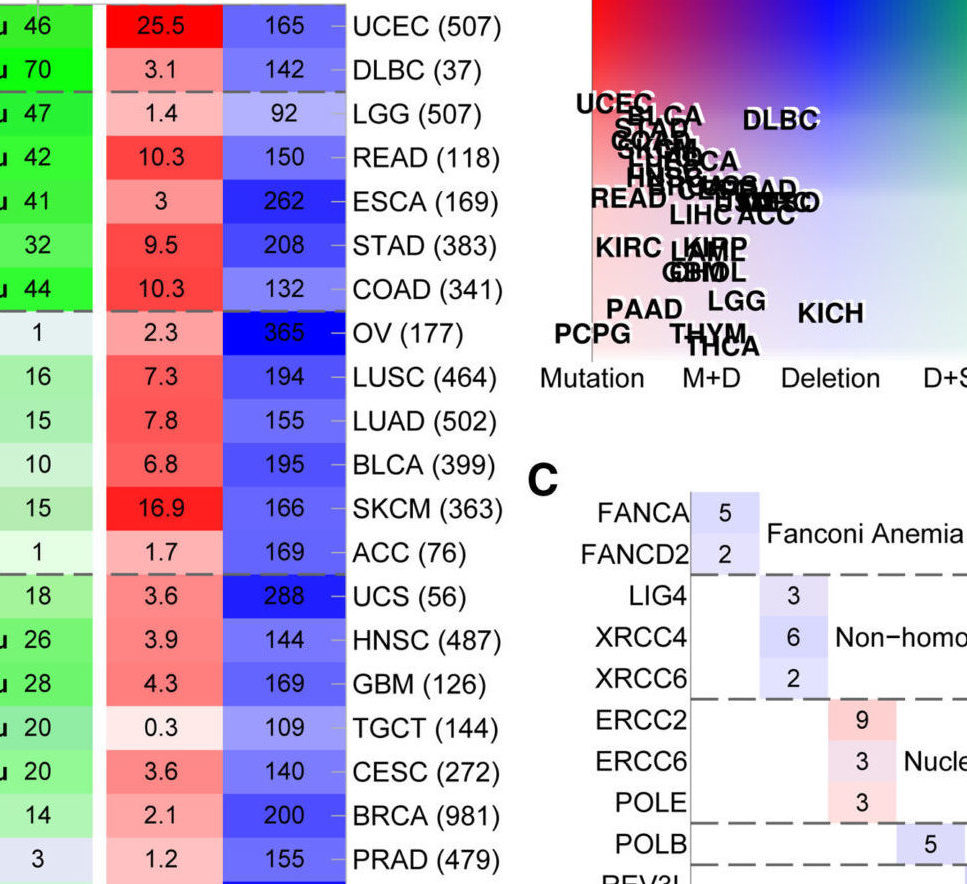
T. Knijnenburg, L. Wang, M. Zimmermann, N. Chambwe, G. Gao, A. Cherniack, H. Fan, H. Shen, G. Way, C. Greene, Y. Liu, R. Akbani, B. Feng, L. Donehower, C. Miller, Y. Shen, M. Karimi, H. Chen, P. Kim, P. Jia, E. Shinbrot, S. Zhang, J. Liu, H. Hu, M. Bailey, C. Yau, D. Wolf, Z. Zhao, J. Weinstein, L. Li, L. Ding, G. Mills, P. Laird, D. Wheeler, I. Shmulevich, Cancer Genome Atlas Research Network, R. Monnat, Y. Xiao, C. Wang, “Genomic and Molecular Landscape of DNA Damage Repair Deficiency across The Cancer Genome Atlas“, Cell Reports, Vol. 23, No. 1, pp. 239-254.e6, 2018.
Q. Gao, W. Liang, S. Foltz, G. Mutharasu, R. Jayasinghe, S. Cao, W. Liao, S. Reynolds, M. Wyczalkowski, L. Yao, L. Yu, S. Sun, K. Chen, A. Lazar, R. Fields, M. Wendl, B. Van Tine, R. Vij, F. Chen, M. Nykter, I. Shmulevich, L. Ding, Cancer Genome Atlas Research Network, “Driver Fusions and Their Implications in the Development and Treatment of Human Cancers“, Cell Reports, Vol. 23, No. 1, pp. 227-238.e3, 2018.
J. Saltz, R. Gupta, L. Hou, T. Kurc, P. Singh, V. Nguyen, D. Samaras, K. Shroyer, T. Zhao, R. Batiste, J. Van Arnam, Cancer Genome Atlas Research Network, I. Shmulevich, A. Rao, A. Lazar, A. Sharma, V. Thorsson, “Spatial Organization and Molecular Correlation of Tumor-Infiltrating Lymphocytes Using Deep Learning on Pathology Images“, Cell Reports, Vol. 23, No. 1, pp. 181-193.e7, 2018.
The Cancer Genome Atlas Research Network, “Comprehensive and Integrated Genomic Characterization of Adult Soft Tissue Sarcomas,” Cell, Vol. 171, No. 4, pp. 950–965.e28, 2017.
The Cancer Genome Atlas Research Network, “Comprehensive and Integrative Genomic Characterization of Hepatocellular Carcinoma,” Cell, Vol. 169, No. 7, pp. 1327–1341.e23, 2017.
V. Kytola, U. Topaloglu, L. D. Miller, R. L. Bitting, M. M. Goodman, J. D. Agostino RB, R. J. Desnoyers, C. Albright, G. Yacoub, S. A. Qasem, B. DeYoung, V. Thorsson, I. Shmulevich, M. Yang, A. Shcherban, M. Pagni, L. Liu, M. Nykter, K. Chen, G. A. Hawkins, S. C. Grant, W. J. Petty, A. T. Alistar, E. A. Levine, E. D. Staren, C. D. Langefeld, V. Miller, G. Singal, R. M. Petro, M. Robinson, W. Blackstock, B. L. Powell, L. I. Wagner, K. L. Foley, E. Abraham, B. Pasche, and W. Zhang, “Mutational Landscapes of Smoking-Related Cancers in Caucasians and African Americans: Precision Oncology Perspectives at Wake Forest Baptist Comprehensive Cancer Center,” Theranostics, Vol. 7, pp. 2914-2923, 2017.
W. Poole, K. Leinonen, I. Shmulevich, T. Knijnenburg, B. Bernard, “Multiscale mutation clustering algorithm identifies pan-cancer mutational clusters associated with pathway-level changes in gene expression,” PLoS Computational Biology, Vol. 13, No. 2, e1005347, 2017.
The Cancer Genome Atlas Research Network, “Comprehensive Molecular Characterization of Pheochromocytoma and Paraganglioma,” Cancer Cell, Vol. 31, No. 2, pp.181-193, 2017.
The Cancer Genome Atlas Research Network, “Integrated genomic characterization of oesophageal carcinoma,” Nature, Vol. 541, pp. 169–175, 2017.
The Cancer Genome Atlas Research Network, “Integrated genomic and molecular characterization of cervical cancer,” Nature, Vol. 543, pp. 387-384, 2017.
Y. Sun, P. Ji, T. Chen, X. Zhou, D. Yang, Y. Guo, Y. Liu, L. Hu, D. Xia, Y. Liu, A. S. Multani, I. Shmulevich, R. Kucherlapati, S. Kopetz, A. K. Sood, S. R. Hamilton, B. Sun, and W. Zhang, “MIIP haploinsufficiency induces chromosomal instability and promotes tumour progression in colorectal cancer,” The Journal of Pathology, Vol. 241, No. 1, pp. 67–79, 2017.
T. A. Knijnenburg, G. W. Klau, F. Iorio, M. J. Garnett, U. McDermott, I. Shmulevich, L. F. A. Wessels, “Logic models to predict continuous outputs based on binary inputs with an application to personalized cancer therapy,” Scientific Reports, Vol. 6, 36812, 2016.
The Cancer Genome Atlas Research Network, “Comprehensive Pan-Genomic Characterization of Adrenocortical Carcinoma,” Cancer Cell, Vol. 29, No. 5, pp. 723-736, 2016.
T. Knijnenburg, T. Bismeijer, L. Wessels, I. Shmulevich, “A multilevel pan-cancer map links gene mutations to cancer hallmarks,” Chinese Journal of Cancer, Vol. 34, 48, 2015.
The Cancer Genome Atlas Research Network, “Genomic Classification of Cutaneous Melanoma,” Cell, Vol. 161, No. 7, pp. 1681–1696, 2015.
The Cancer Genome Atlas Research Network, “The Molecular Taxonomy of Primary Prostate Cancer,” Cell, Vol. 163 , No. 4 , pp. 1011-1025, 2015.
The Cancer Genome Atlas Research Network, “Comprehensive, Integrative Genomic Analysis of Diffuse Lower-Grade Gliomas,” The New England Journal of Medicine, Vol. 372, pp. 2481-2498, 2015.
Y. Liu, M. Yasukawa, K. Chen, L. Hu, R. R. Broaddus, L. Ding, E. R. Mardis, P. Spellman, D. A. Levine, G. B. Mills, I. Shmulevich, A. K. Sood, W. Zhang, “Association of Somatic Mutations of ADAMTS Genes With Chemotherapy Sensitivity and Survival in High-Grade Serous Ovarian Carcinoma,” JAMA Oncology, Vol. 1, No. 4, pp. 486-494, 2015.
Y. Sun, F. Guo, M. Bagnoli, F.-X. Xue, B.-C. Sun, I. Shmulevich, D. Mezzanzanica, K.-X. Chen, A. Sood, D. Yang, W. Zhang, “Key nodes of a microRNA network associated with the integrated mesenchymal subtype of high-grade serous ovarian cancer,” Chinese Journal of Cancer, Vol. 34, No. 1, pp. 28-40, 2015.
G. Liu, D. Yang, R. Rupaimoole, C. V. Pecot, Y. Sun, L. S. Mangala, X. Li, P. Ji, D. Cogdell, L. Hu, Y. Wang, C. Rodriguez-Aguayo, G. Lopez-Berestein, I. Shmulevich, L. De Cecco, K. Chen, D. Mezzanzanica, F. Xue, A. K. Sood, W. Zhang, “Augmentation of Response to Chemotherapy by microRNA-506 Through Regulation of RAD51 in Serous Ovarian Cancers,” Journal of the National Cancer Institute, Vol. 107, No. 7, djv108, 2015.
K. M. Turner, Y. Sun, P. Ji, K. J. Granberg, B. Bernard, L. Hu, D. E. Cogdell, X. Zhou, O. Yli-Harja, M. Nykter, I. Shmulevich, W. K. Yung, G. N. Fuller, W. Zhang, “Genomically amplified Akt3 activates DNA repair pathway and promotes glioma progression,” Proceedings of the National Academy of Sciences of the USA, Vol. 112, No. 11, pp. 3421-3426, 2015.
The Cancer Genome Atlas Research Network, “Integrated Genomic Characterization of Papillary Thyroid Carcinoma,” Cell, Vol. 159, No. 3, pp. 676–690, 2014.
The Cancer Genome Atlas Research Network, “Comprehensive Molecular Characterization of Gastric Adenocarcinoma,” Nature, Vol. 513, pp. 202–209, 2014.
C. J. Kemp, J. M. Moore, R. Moser, B. Bernard, M. Teater, L. E. Smith, N. Rabaia, K. E. Gurley, J. Guinney, S. E. Busch, R. Shaknovich, V. V. Lobanenkov, D. Liggitt, I. Shmulevich, A. Melnick, G. N. Filippova, “CTCF haploinsufficiency destabilizes DNA methylation and predisposes to cancer,” Cell Reports, Vol. 7, No. 4, pp. 1020-1029, 2014.
X. Li, Y. Liu, K. J. Granberg, Q. Wang, L. M. Moore, P. Ji, J. Gumin, E. P. Sulman, G. A. Calin, H. Haapasalo, M. Nykter, I. Shmulevich, G. N. Fuller, F. F. Lang, W. Zhang, “Two mature products of MIR-491 coordinate to suppress key cancer hallmarks in glioblastoma,” Oncogene, Vol. 34, pp. 1619-1628, 2014.
The Cancer Genome Atlas Research Network, “Comprehensive molecular characterization of urothelial bladder carcinoma,” Nature, Vol. 507, pp. 315–322, 2014.
G. Liu, Y. Sun, P. Ji, X. Li, D. Cogdell, D. Yang, B. C. Parker-Kerrigan, I. Shmulevich, K. Chen, A. K. Sood, F. Xue, W. Zhang, “MiR-506 Suppresses Proliferation and Induces Senescence by Directly Targeting the CDK4/6-FOXM1 Axis in Ovarian Cancer,” The Journal of Pathology, Vol. 233, No. 3, pp. 308-318, 2014.
Y. Liu , L. Patel , G. B. Mills, K. H. Lu , A. K. Sood , L. Ding, R. Kucherlapati , E. Mardis, D. A. Levine, I. Shmulevich, R. R. Broaddus, W. Zhang, “Clinical Significance of CTNNB1 Mutation and Wnt Pathway Activation in Endometrioid Endometrial Carcinoma,” Journal of the National Cancer Institute, Vol. 106, No. 9, 2014.
F. Guo, B. C. Parker, D. Yang, L. Hu, I. Shmulevich, A. K. Sood, F. Xue, W. Zhang, “Post-Transcriptional Regulatory Network of Epithelial-to-Mesenchymal and Mesenchymal-to-Epithelial Transitions,” Journal of Hematology & Oncology, Vol. 7, No. 19, 2014.
J. Ashworth, B. Bernard, S. Reynolds, C. L. Plaisier, I. Shmulevich, N. S. Baliga, “Structure-based predictions broadly link transcription factor mutations to gene expression changes in cancers,” Nucleic Acids Research, Vol. 42, No. 21, pp. 12973-12983, 2014.
C. W. Brennan, R.G.W. Verhaak, A. McKenna, B. Campos, H. Noushmehr, S. R. Salama, S. Zheng, D. Chakravarty, J. Z. Sanborn, S. H. Berman, R. Beroukhim, B. Bernard, C.J. Wu, G. Genovese, I. Shmulevich, J. Barnholtz-Sloan, L. Zou, R. Vegesna, S. A. Shukla, G. Ciriello, W.K. Yung, W. Zhang, C. Sougnez, T. Mikkelsen, K. Aldape, D. D. Bigner, E. G. Van Meir, M. Prados, A. Sloan, K. L. Black, J. Eschbacher, G. Finocchiaro, W. Friedman, D. W. Andrews, A. Guha, M. Iacocca, B. P. O’Neill, G. Foltz, J. Myers, D. J. Weisenberger, R. Penny, R. Kucherlapati, C. M. Perou, D. N. Hayes, R. Gibbs, M. Marra, G. B. Mills, E. Lander, P. Spellman, R. Wilson, C. Sander, J. Weinstein, M. Meyerson, S. Gabriel, P. W. Laird, D. Haussler, G. Getz, L. Chin, TCGA Research Network, “The Somatic Genomic Landscape of Glioblastoma,” Cell, Vol. 155, No. 2, pp. 462-477, 2013.
The Cancer Genome Atlas Research Network, John N Weinstein, Eric A Collisson, Gordon B Mills, Kenna R Mills Shaw, Brad A Ozenberger, Kyle Ellrott, Ilya Shmulevich, Chris Sander, Joshua M Stuart, “The Cancer Genome Atlas Pan-Cancer analysis project,” Nature Genetics, Vol. 45, No. 10, pp. 1113-1120, 2013.
The Cancer Genome Atlas Research Network, “Integrated genomic characterization of endometrial carcinoma,” Nature, Vol. 497, pp. 67–73, 2013.
D. Yang, Y. Sun, L. Hu, H. Zheng, P. Ji, C. V. Pecot, Y. Zhao, S. Reynolds, H. Cheng, R. Rupaimoole, D. Cogdell, M. Nykter, R. Broaddus, C. Rodriguez-Aguayo, G. Lopez-Berestein, J. Liu, I. Shmulevich, A. K. Sood, K. Chen, W. Zhang, “Integrated Analyses Identify a Master MicroRNA Regulatory Network for the Mesenchymal Subtype in Serous Ovarian Cancer,” Cancer Cell, Vol. 23, No. 2, pp. 186-199, 2013.
L. M. Moore, V. Kivinen, Y. Liu, M. Annala, D. Cogdell, X. Liu, C.G. Liu, R. Sawaya, O. Yli-Harja, I. Shmulevich, G.N. Fuller, W. Zhang, M. Nykter, “Transcriptome and small RNA deep sequencing reveals deregulation of miRNA biogenesis in human glioma,” The Journal of Pathology, Vol. 229, No. 3, pp. 449-459, 2013.
The Cancer Genome Atlas Network, “Comprehensive Molecular Portraits of Human Breast Tumors,” Nature, Vol. 490, No. 7418, pp. 61–70, 2012.
The Cancer Genome Atlas Network, “Comprehensive Molecular Characterization of Human Colon and Rectal Cancer,” Nature, Vol. 487, No. 7407, pp. 330-337, 2012.
Y. Liu, Y. Sun, R. Broaddus, J. Liu, A. K. Sood, I. Shmulevich, Wei Zhang, “Integrated Analysis of Gene Expression and Tumor Nuclear Image Profiles Associated with Chemotherapy Response in Serous Ovarian Carcinoma,” PLoS ONE, Vol. 7, No. 5, e36383, 2012.
G. Liu, D. Yang, Y. Sun, I. Shmulevich, F. Xue, A.K. Sood, W. Zhang, “Differing clinical impact of BRCA1 and BRCA2 mutations in serous ovarian cancer,” Pharmacogenomics, Vol. 13, No. 13, pp. 1523-1535, 2012.
D. Yang, S. Khan, Y. Sun, K. Hess, I. Shmulevich, A. K. Sood, W. Zhang, “Association of BRCA1 and BRCA2 Mutations With Survival, Chemotherapy Sensitivity, and Gene Mutator Phenotype in Patients With Ovarian Cancer,” JAMA, Vol. 306, No. 14, pp. 1557-1565, 2011.
D. Yang, A. Ylipää, J. Yang, K. Hunt, R. Pollock, J. Trent, O. Yli-Harja, I. Shmulevich, M. Nykter, W. Zhang, “An integrated study of aberrant gene copy number and gene expression in GIST and LMS,” Technology in Cancer Research and Treatment, Vol. 9, No. 2, pp. 171-178, 2010.
L. Hu, W. Hittelman, T. Lu, P. Ji, R. Arlinghaus, I. Shmulevich, S. R. Hamilton, W. Zhang, “NGAL decreases E-cadherin-mediated cell-cell adhesion and increases cell motility and invasion through Rac1 in colon carcinoma cells,” Laboratory Investigation, Vol. 89, pp. 531–548, 2009.
N. D. Price, J. Trent, A. K. El-Naggar, D. Cogdell, E. Taylor, K. K. Hunt, R. E. Pollock, L. Hood, I. Shmulevich, W. Zhang, “Highly accurate two-gene classifier for differentiating gastrointestinal stromal tumors and leiomyosarcomas,” Proceedings of the National Academy of Sciences of the USA, Vol. 104, No. 9, pp- 3414-3419, 2007.
R. Jiang, C. Mircean, I. Shmulevich, D. Cogdell, Y. Jia, I. Tabus, K. Aldape, R. Sawaya, J. M. Bruner, G. N. Fuller, W. Zhang, “Pathway alterations during glioma progression revealed by reverse phase protein lysate arrays,” Proteomics, Vol. 6, pp. 2964-2971, 2006.
M. Nykter, K. K. Hunt, R. E. Pollock, A. K. El-Naggar, E. Taylor, I. Shmulevich, O. Yli-Harja, W. Zhang, “Unsupervised analysis uncovers changes in histopathologic diagnosis in supervised genomic studies,” Technology in Cancer Research and Treatment, Vol. 5, No. 2, pp. 177-182, 2006.
G. N. Fuller, C. Mircean, I. Tabus, E. Taylor, R. Sawaya, J. M. Bruner, I. Shmulevich, W. Zhang, “Molecular voting for glioma classification reflecting heterogeneity in the continuum of cancer progression,” Oncology Reports, Vol. 14, pp. 651-656, 2005.
E. Lee, C. Mircean, I. Shmulevich, H. Wang, J. Liu, A. Niemistö, J. Kavanagh, JH Lee, W. Zhang, “Insulin-like growth factor binding protein 2 promotes ovarian cancer cell invasion,” Molecular Cancer, Vol. 4, No. 7, 2005.
H. Lähdesmäki, X. Hao, B. Sun, L. Hu, O. Yli-Harja, I. Shmulevich, and W. Zhang, “Distinguishing key biological pathways between primary breast cancers and their lymph node metastases by gene function-based clustering analysis,” International Journal of Oncology, Vol. 24, No. 6, pp. 1589–1596, June 2004.
X. Hao, B. Sun, L. Hu, H. Lähdesmäki, V. Dunmire, Y. Feng, S.-W. Zhang, H. Wang, C. Wu, H. Wang, G. N. Fuller, W. F. Symmans, I. Shmulevich, and W. Zhang, “Differential gene and protein expression in primary breast malignancies and their lymph node metastases as revealed by combined cDNA microarray and tissue microarray analysis,” Cancer, Vol. 100, No. 6, pp. 1110-1122, 2004.
C. Mircean, I. Tabus, T. Kobayashi, M. Yamaguchi, H. Shiku, I. Shmulevich, W. Zhang, “Pathway Analysis of Informative Genes from Microarray Data Reveals that Metabolism and Signal Transduction Genes Distinguish Different Subtypes of Lymphomas,” International Journal of Oncology, Vol. 24, No. 3, pp. 497-504, 2004.
J. Morikawa, H. Li, S. Kim, K. Nishi, S. Ueno, E. Suh, E. Dougherty, I. Shmulevich, H. Shiku, W. Zhang, and T. Kobayashi, “Identification of signature genes by microarray for acute myeloid leukemia without maturation and acute promyelocytic leukemia with t(15;17)(q22;q12)(PML/RARa),” International Journal of Oncology, Vol. 23, No. 3, pp. 617-625, 2003.
T. Kobayashi, M. Yamaguchi, S. Kim, J. Morikawa, S. Ogawa, S. Ueno, E. Suh, E. Dougherty, I. Shmulevich, H. Shiku, W. Zhang, “Microarray Reveals Differences in Both Tumors and Vascular Specific Gene Expression in De Novo CD5+ and CD5- Diffuse Large B-cell Lymphomas,” Cancer Research, vol. 63, pp. 60-66, January 2003.
S. Kim, E. R. Dougherty, I. Shmulevich, K. R. Hess, S. R. Hamilton, J. M. Trent, G. N. Fuller, W. Zhang, “Identification of Combination Gene Sets for Glioma Classification,” Molecular Cancer Therapeutics, vol. 1, pp. 1229-1236, November 2002.
I. Shmulevich, K. Hunt, A. El-Naggar, E. Taylor, L. Ramdas, P. Labordé, K. R. Hess, R. Pollock, W. Zhang, “Tumor Specific Gene Expression Profiles in Human Leiomyosarcoma: an Evaluation of Intra-Tumor Heterogeneity,” Cancer, Vol. 94, No. 7, pp. 2069-2075, 2002. [/su_expand]
Featured Projects
-
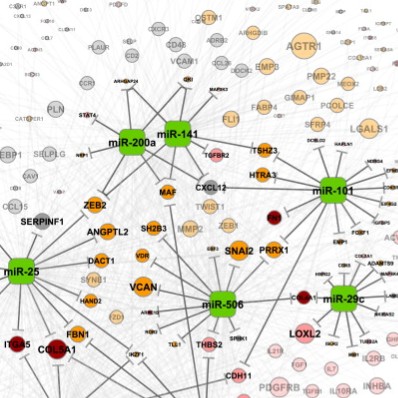
The study of microRNAs in cancer
MicroRNAs (miRNAs) are small non-coding RNAs that can regulate gene expression and that are frequently altered in human cancers. Computational approaches can help us understand how miRNAs are dysregulated and potentially offer new therapies.
-

Cancer Target Discovery and Development
The goal of the CTD2 Network is to understand how tumor heterogeneity leads to drug resistance in order to develop optimal combinations of chemotherapy or small molecules in combination with immunotherapy.
-
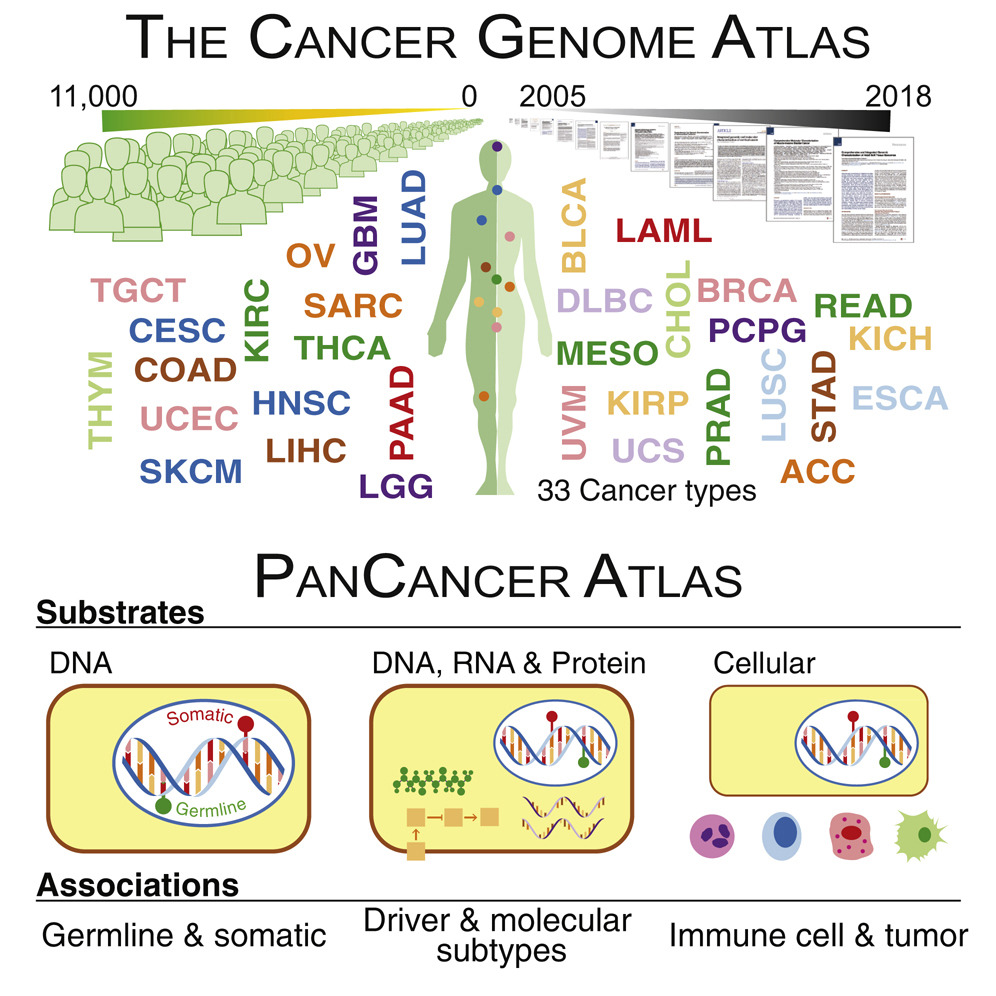
Pan-Cancer Atlas
From the analysis of over 11,000 tumors from 33 of the most prevalent forms of cancer, the Pan-Cancer Atlas provides a uniquely comprehensive, in-depth, and interconnected understanding of how, where, and why tumors arise in humans.
-
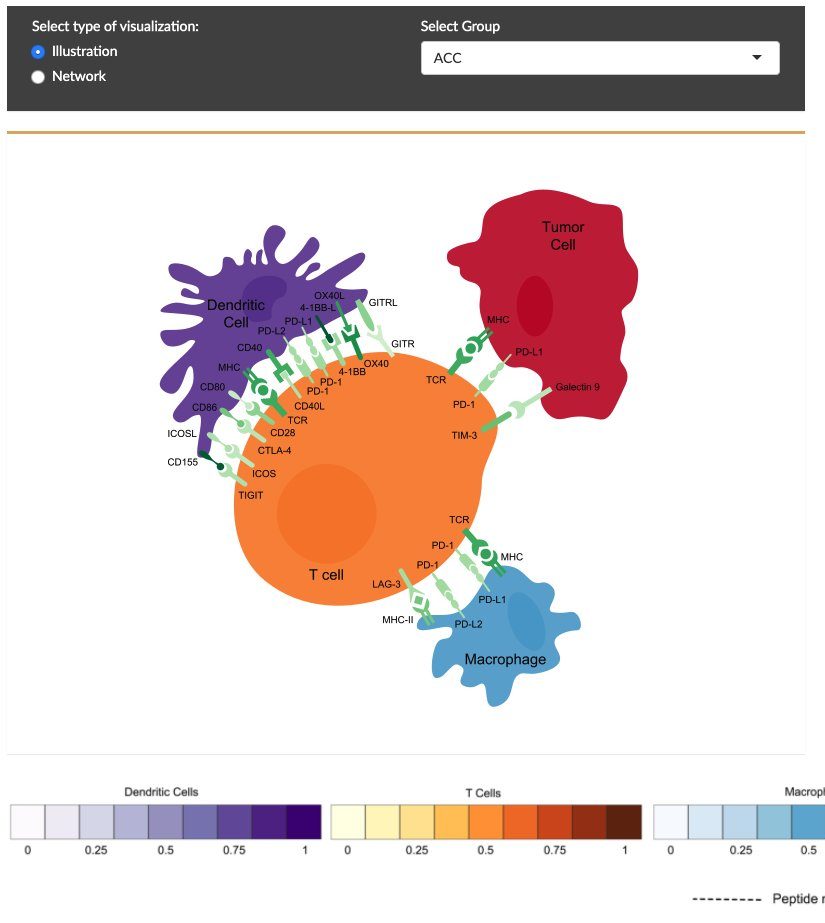
CRI iAtlas
The Cancer Research Institute (CRI) iAtlas is an interactive web-based platform and set of analytic tools for studying interactions between tumors and the immune microenvironment.
-
Human Tumor Atlas Network
Data Coordinating Center for the Human Tumor Atlas Network
-
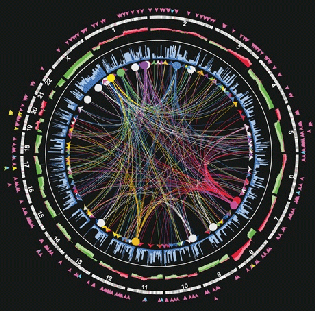
TCGA: Genome Data Analysis Center
Our group had the privilege of being part of The Cancer Genome Atlas (TCGA), a landmark cancer genomics program that molecularly characterized over 20,000 primary cancer and matched normal samples spanning 33 cancer types.
-

The ISB Cancer Genomics Cloud
The ISB Cancer Genomics Cloud (ISB-CGC) is democratizing access to NCI Cancer Data and coupling it with unprecedented computational power to allow researchers to explore and analyze this vast data-space.


 shmulevich.isbscience.org/research/cancer-studies/
shmulevich.isbscience.org/research/cancer-studies/